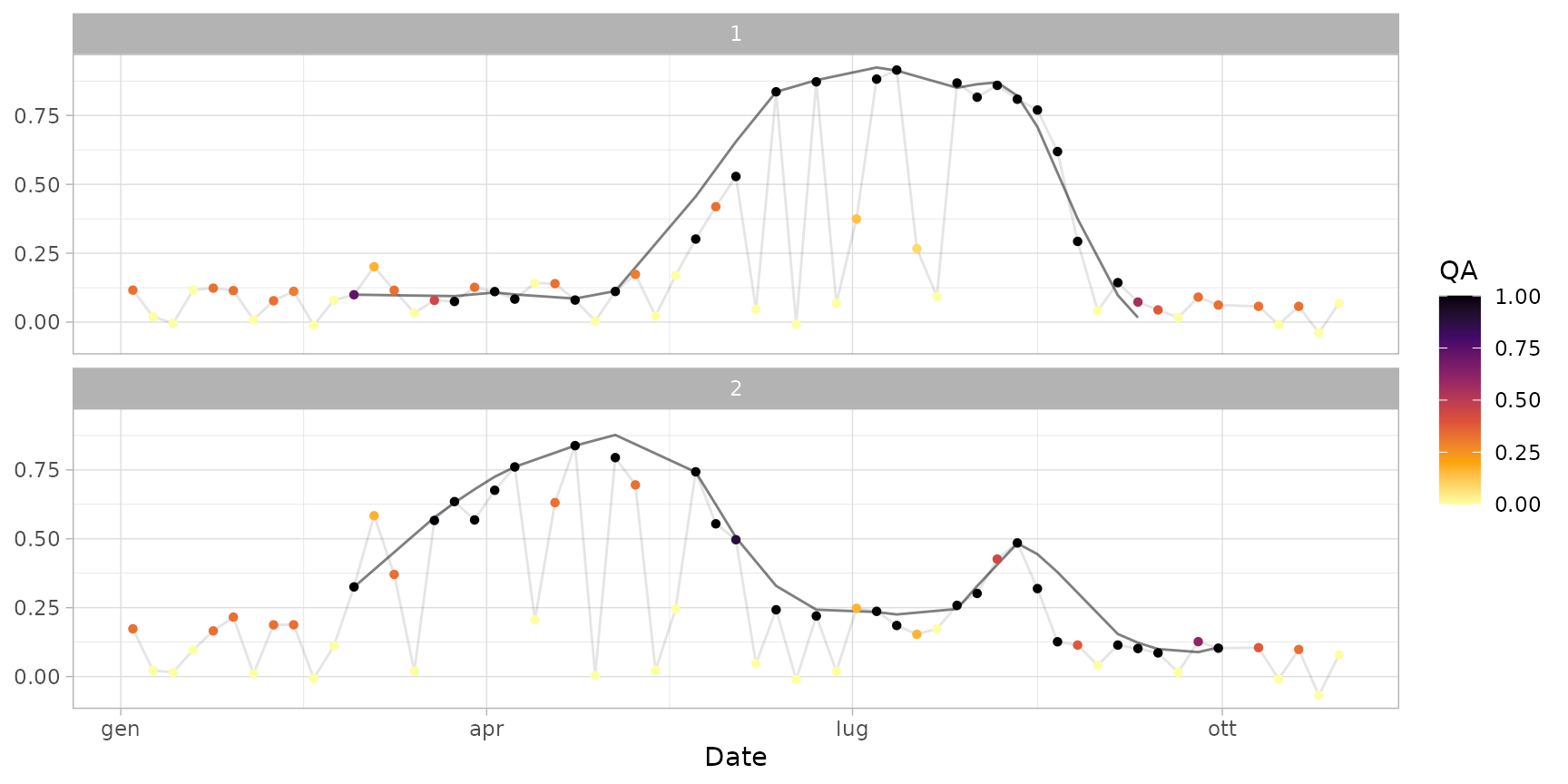
Smooth time series extracted with function extract_s2ts()
using a Savitzky-Golay filter.
Quality flags associated to the time series are used to obtain better
results.
Argument values should be manually set to obtain a correct smoothing.
smooth_s2ts(
ts,
min_qa = 0.2,
noise_dir = "low",
spike = 0.25,
spike_window = 5,
sg_daywindow = 15,
sg_polynom = 2,
sg_n = 3,
max_extrapolation = 0.1
)Arguments
- ts
Time series in
s2tsformat (generated usingextract_s2ts()).- min_qa
(optional) minimum 0-1 quality value (points with
qa < min_qaare not used, whileqavalues in the rangemin_qato 1 are reshaped into the range 0 to 1). Default is 0.5.- noise_dir
Direction of points generally containing noise. If
"low", higher values are generally maintained (this is the case of most vegetation indices like NDVI); if"high", lower values are generally maintained; if"undefined"(default), no assumptions are done.- spike
Relative "y" difference for spike determination (default is 0.25). Set to NA to skip spike removal.
- spike_window
Maximum number of values for spike identification (it must be an odd number). Default is 3.
- sg_daywindow
Half-size of the time window to be used to interpolate
- sg_polynom
Argument
polynomof functionw_savgol().- sg_n
(optional) Positive integer: number of applications of the Savitzky-Golay filter. The minimum value is 1 (a single application using the weights included in
ts). Within each additional application, the weights included intsare multiplicated for the relative ranks of the difference between original and smoothed values (ifnoise_dir = "low") or between smoothed and original values (ifnoise_dir = "high"). Ifnoise_dir = "undefined", this argument is coerced to 1.- max_extrapolation
(optional) Numeric: maximum allowed extrapolation out of original range (relative value). Default is 0.1 (+10%). Set to Inf in order not to set any constraint.
Value
The output time series in s2ts format.
Examples
# Load input data
data("ts_raw")
# Smoothing time series using default parameters
ts_smoothed <- smooth_s2ts(ts_raw)
print(ts_smoothed, topn = 5) # standard print
#> A smoothed s2ts time series with 60 dates and 2 IDs.
#> Date Orbit Sensor 1 2
#> 1: 2020-01-04 022 2B 0.11606015 ○ 0.1740709 ○
#> 2: 2020-01-09 022 2A NA ○ NA ○
#> 3: 2020-01-14 022 2B NA ○ NA ○
#> 4: 2020-01-19 022 2A NA ○ NA ○
#> 5: 2020-01-24 022 2B 0.12343408 ○ 0.2060596 ○
#> ---
#> 56: 2020-10-10 022 2B 0.05760022 ○ 0.1021135 ○
#> 57: 2020-10-15 022 2A NA ○ NA ○
#> 58: 2020-10-20 022 2B 0.05767390 ○ 0.1016156 ○
#> 59: 2020-10-25 022 2A NA ○ NA ○
#> 60: 2020-10-30 022 2B NA ○ NA ○
#>
#> Quality flags: ● [1] ◕ [0.9,1) ◑ [0.75,0.9) ◔ [0.5,0.75) ○ [0,0.5)
head(as.data.frame(ts_smoothed)) # see content
#> id date orbit sensor value qa rawval
#> 1 1 2020-01-04 022 2B 0.1160602 0.33 0.116133333
#> 2 1 2020-01-09 022 2A NA 0.00 0.020170833
#> 3 1 2020-01-14 022 2B NA 0.00 -0.004372917
#> 4 1 2020-01-19 022 2A NA 0.00 0.116368750
#> 5 1 2020-01-24 022 2B 0.1234341 0.33 0.123387500
#> 6 1 2020-01-29 022 2A 0.1137065 0.33 0.114379167
plot(ts_smoothed)
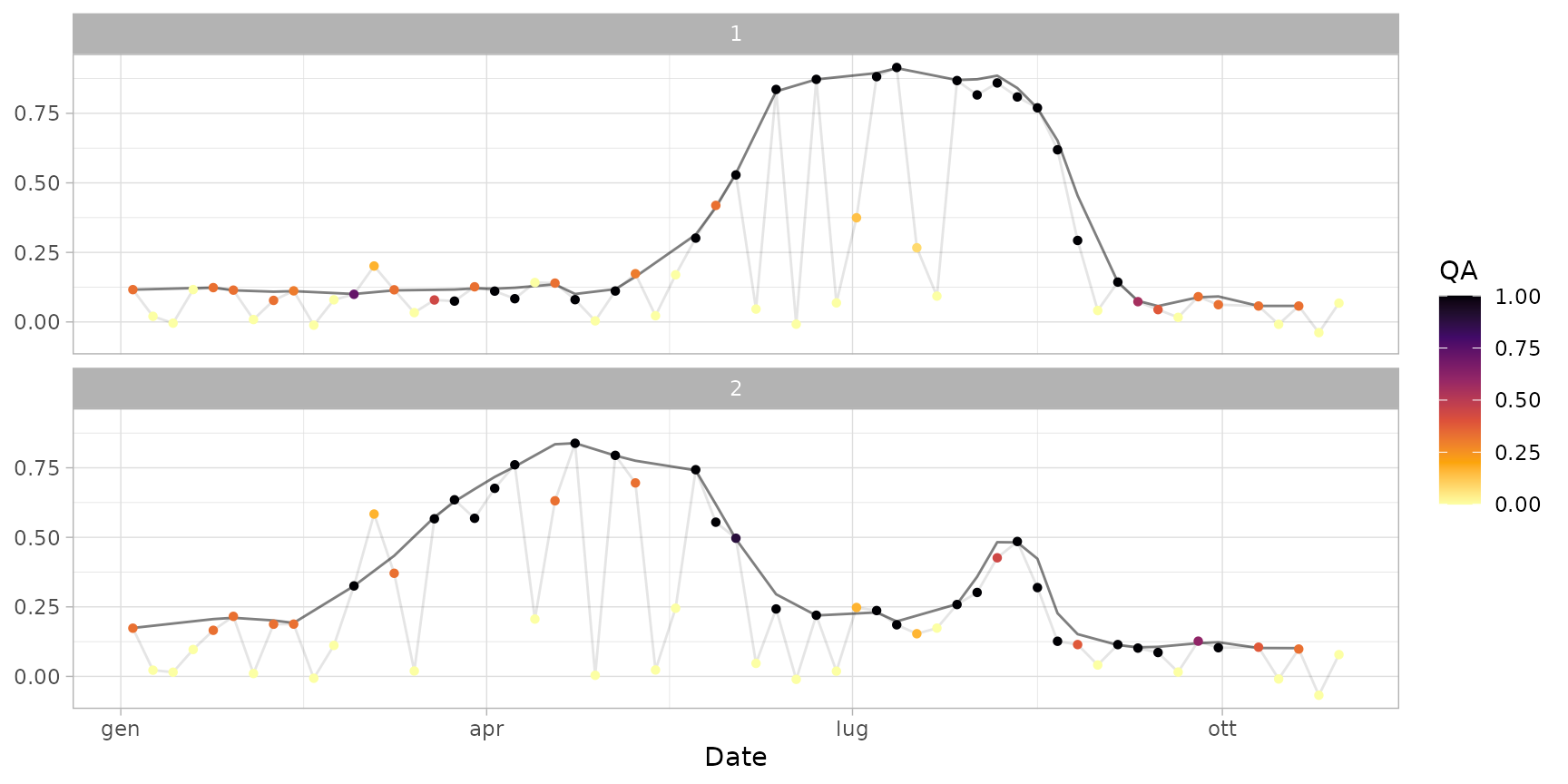 # Apply a more pronounced smoothing
ts_smoothed_2 <- smooth_s2ts(
ts_raw,
min_qa = 0.5, # exclude values with qa < 0.5
sg_daywindow = 30, # larger moving window
sg_polynom = 3, # cubic interpolation
sg_n = 5 # apply the SG filter 5 times
)
plot(ts_smoothed_2)
# Apply a more pronounced smoothing
ts_smoothed_2 <- smooth_s2ts(
ts_raw,
min_qa = 0.5, # exclude values with qa < 0.5
sg_daywindow = 30, # larger moving window
sg_polynom = 3, # cubic interpolation
sg_n = 5 # apply the SG filter 5 times
)
plot(ts_smoothed_2)