Extract phenological metrics from a fitted time series.
extract_pheno(data, method = "trs", trs = 0.5, ...)Arguments
- data
List of fitted time series as generated by function
fit_curve().- method
Thresholding method among
"trs","derivatives","klosterman"and"gu"(seephenopix::PhenoExtract()).- trs
Argument passed to
phenopix::PhenoExtract()ifmethod = "trs". It is valid also formethod = "derivatives"thanks to a{sen2rts}implementation (it will be documented soon). Other methods do not make use of this argument.- ...
Additional arguments passed to
phenopix::PhenoExtract().
Value
A data table with the following fields:
id: the time series ID (sees2ts);year: the year (integer);cycle: the cycle ID (integer);begin,end,maxval: the dates of begin, end and maximum value in the cycle as computed bycut_cycles();other phenology metrics, depending on
method:if
method = "trs"or"derivatives":sos,eos,los,pop,mgs,rsp,rau,peak,msp,mau(seephenopix::PhenoTrs()orphenopix::PhenoDeriv());if
method = "gu":UD,SD,DD,RD,maxline,baseline,prr,psr,plateau.slope(seephenopix::PhenoGu());if
method = "klosterman":Greenup,Maturity,Senescence,Dormancy(seephenopix::PhenoKl()).
Examples
# Load input data
data("cf")
data("ts_filled") # used for plots
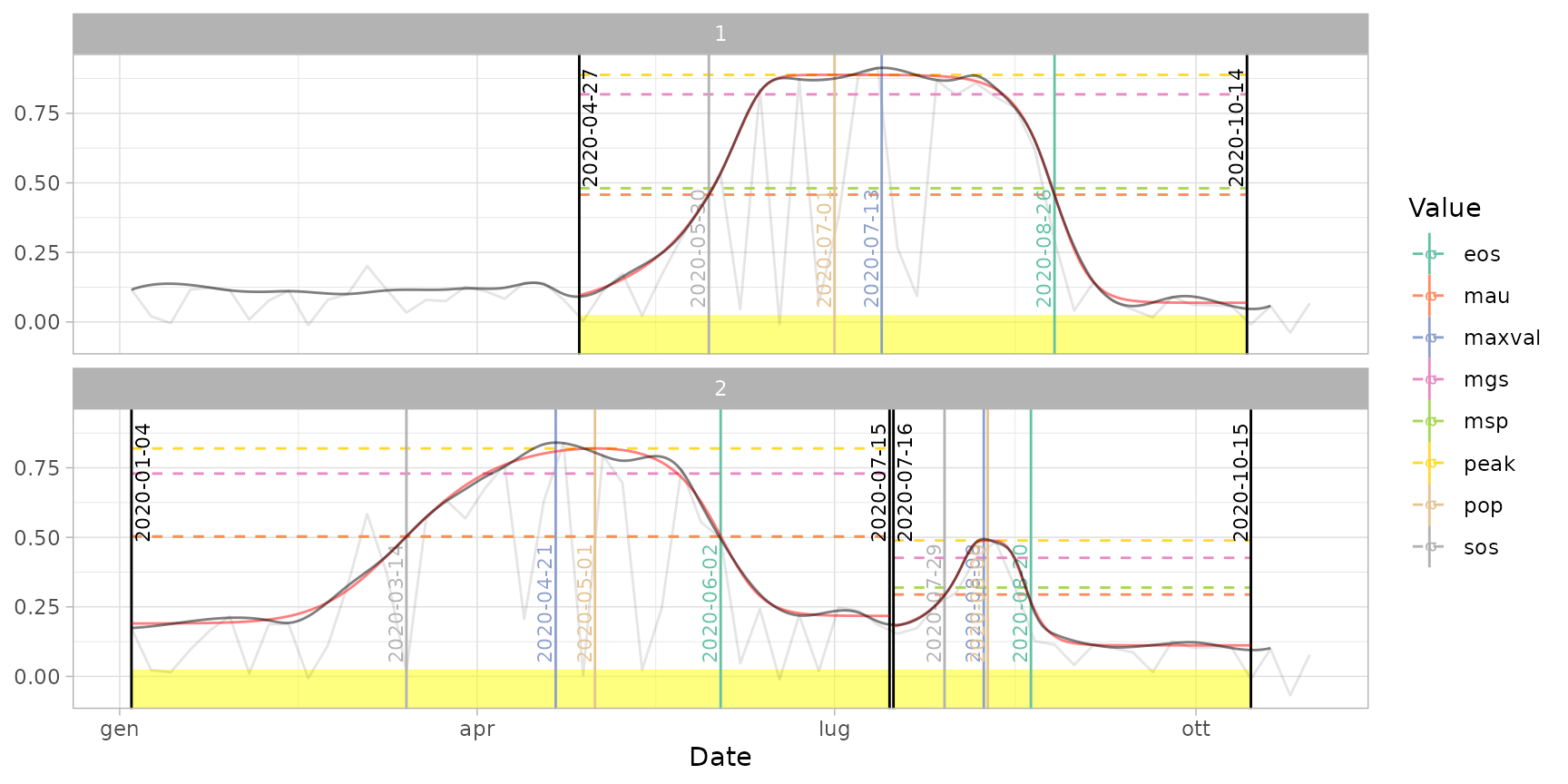
# Default extraction ("trs" method with 50% threshold)
dt_pheno <- extract_pheno(cf)
plot(ts_filled, fitted = cf, pheno = dt_pheno, plot_dates = TRUE)
 # \donttest{
# Customize parameters (e.g. "derivatives" method with 30% threshold)
dt_pheno_2 <- extract_pheno(cf, method = "derivatives", trs = 0.3)
plot(ts_filled, fitted = cf, pheno = dt_pheno_2, plot_dates = TRUE)
# \donttest{
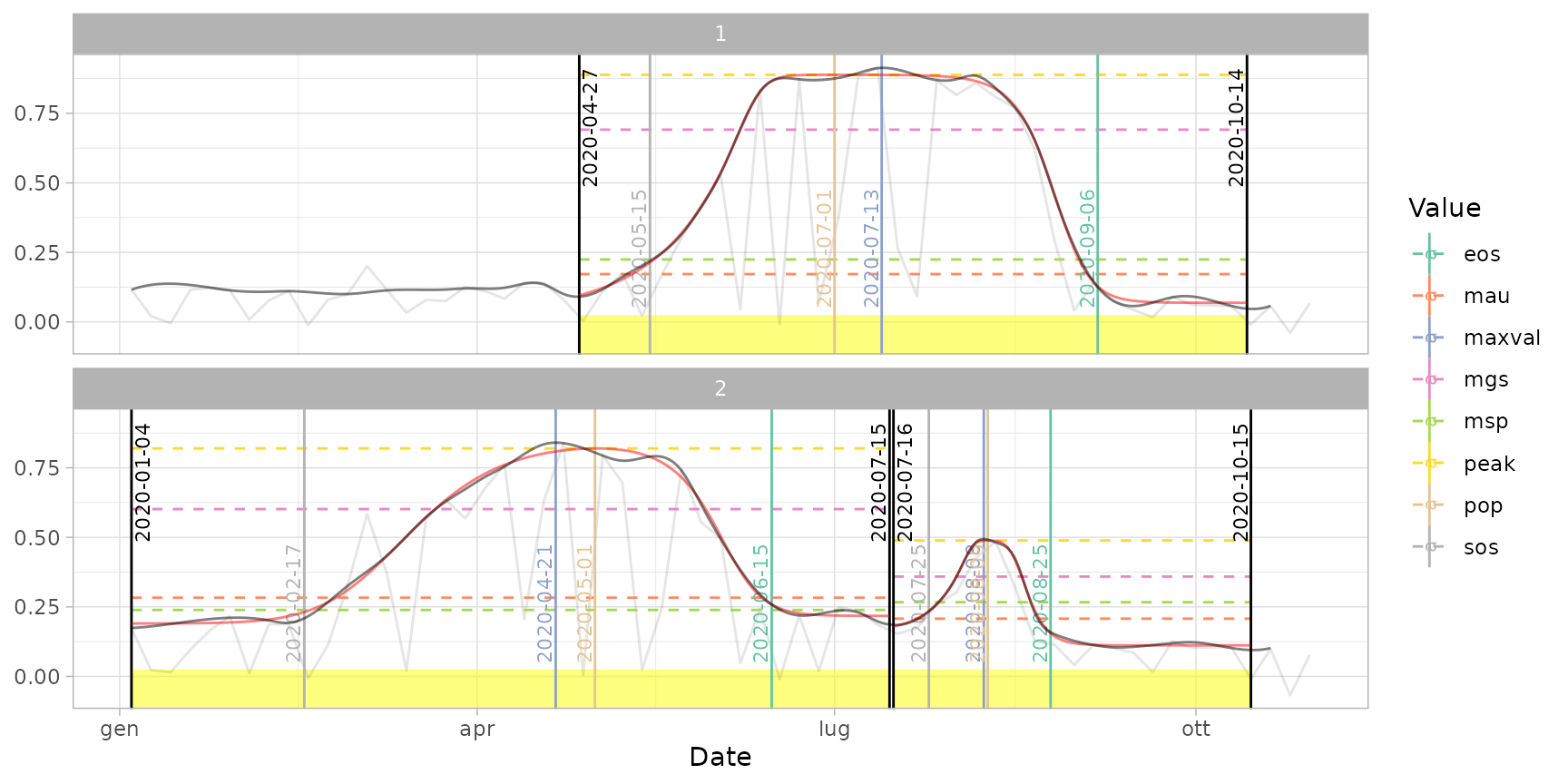
# Customize parameters (e.g. "derivatives" method with 30% threshold)
dt_pheno_2 <- extract_pheno(cf, method = "derivatives", trs = 0.3)
plot(ts_filled, fitted = cf, pheno = dt_pheno_2, plot_dates = TRUE)
 # Other methods: Klosterman
dt_pheno_kl <- extract_pheno(cf, method = "klosterman")
plot(ts_filled, fitted = cf, pheno = dt_pheno_kl, plot_dates = TRUE)
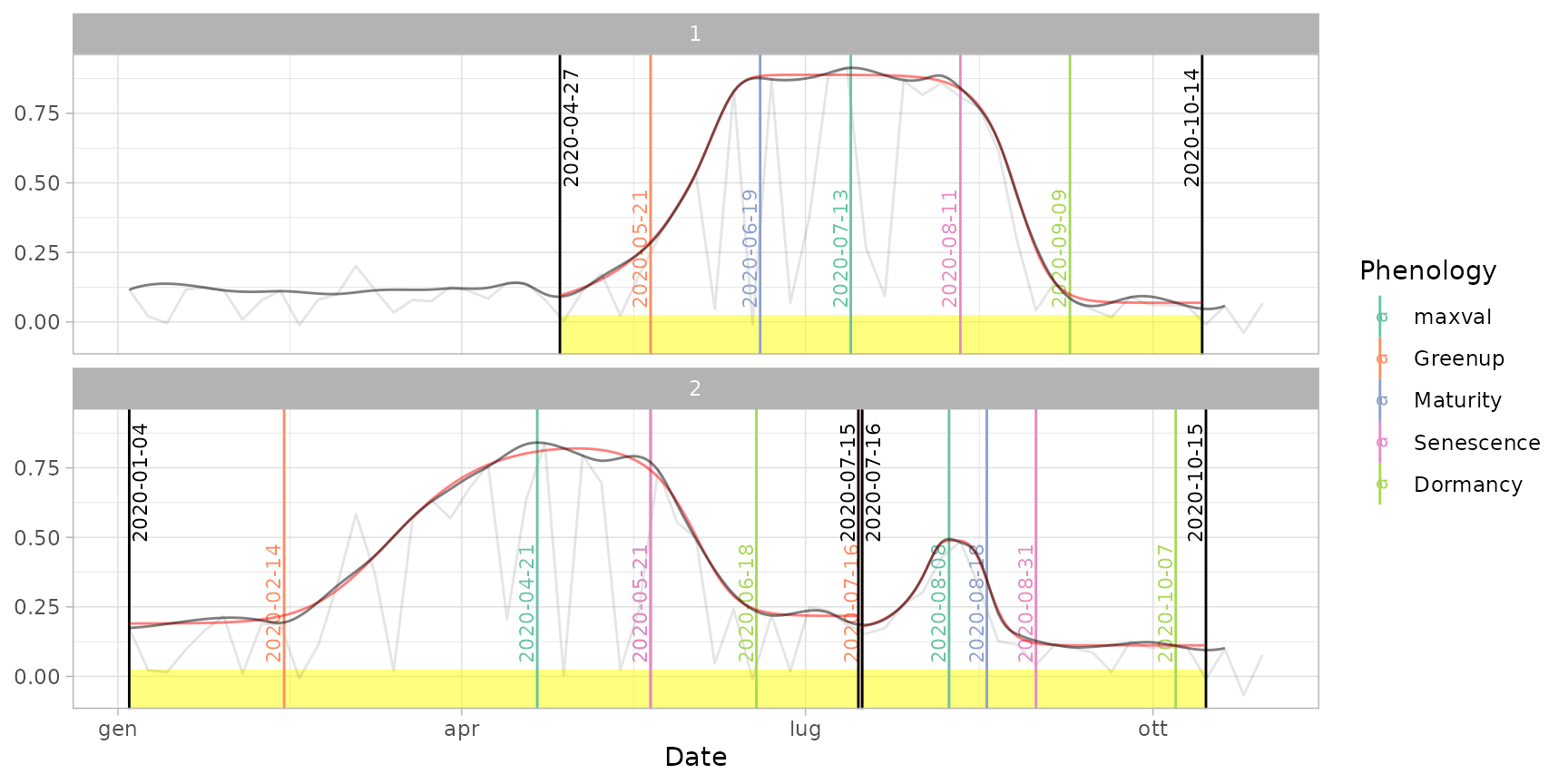
# Other methods: Klosterman
dt_pheno_kl <- extract_pheno(cf, method = "klosterman")
plot(ts_filled, fitted = cf, pheno = dt_pheno_kl, plot_dates = TRUE)
 # Other methods: Gu
dt_pheno_gu <- extract_pheno(cf, method = "gu")
plot(ts_filled, fitted = cf, pheno = dt_pheno_gu, plot_dates = TRUE)
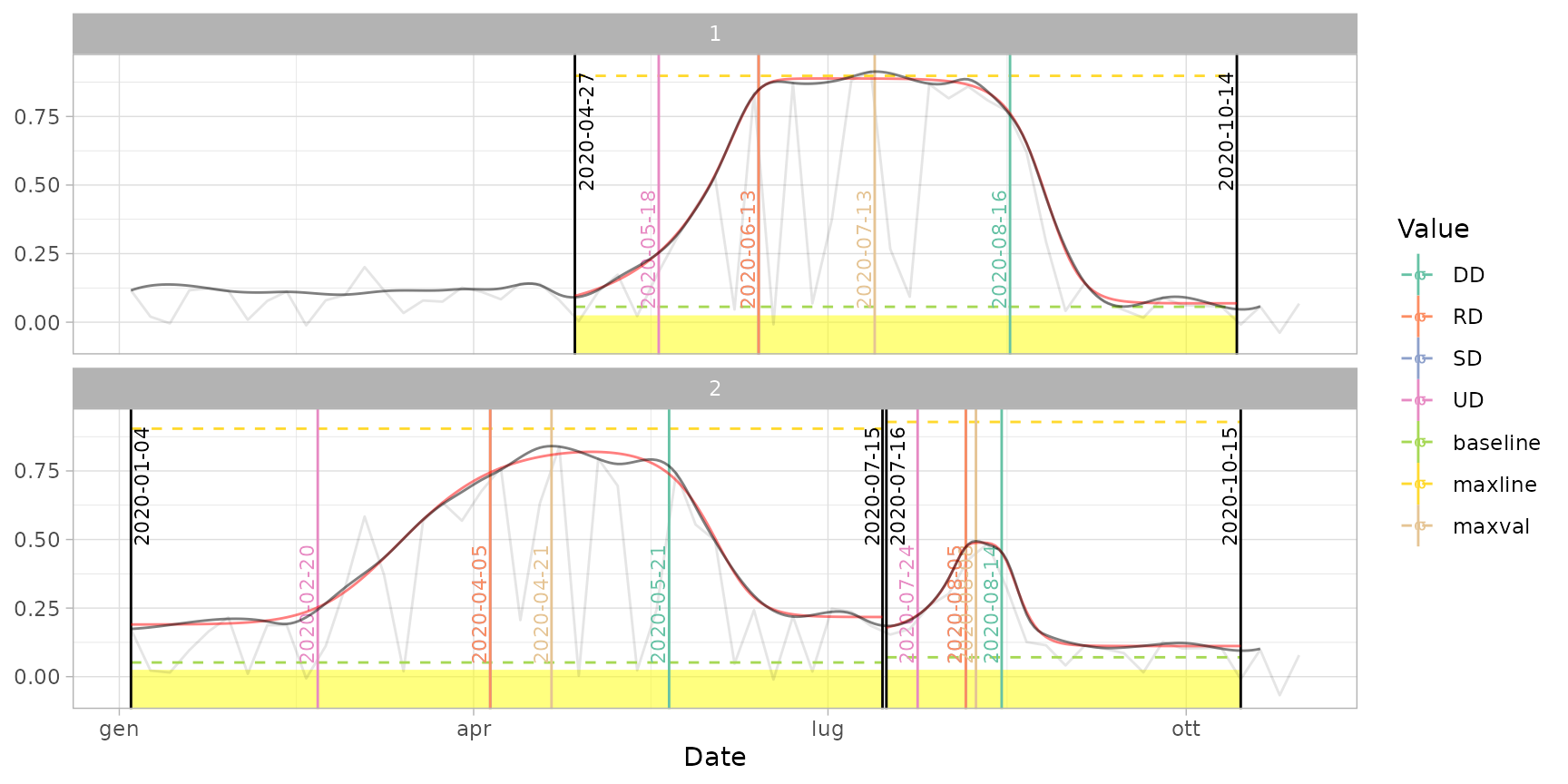
# Other methods: Gu
dt_pheno_gu <- extract_pheno(cf, method = "gu")
plot(ts_filled, fitted = cf, pheno = dt_pheno_gu, plot_dates = TRUE)
 # }
# }